



Next: Alignment using an initial
Up: Worked examples
Previous: Worked examples
Mammalian and Bacterial Serine Proteinases
(This example is discussed in Russell & Barton, (1992).)
Despite a pronounced functional similarity (a highly conserved catalytic
triad), this family of proteins shows little overall sequence similarity.
Indeed, sequence alignment methods generally fail to provide an accurate
alignment of these protein sequences. In situations like these, STAMP
can be used to provide an accuarate alignment of protein sequences based
on a comparison of 3D structure. This can often reveal regions of weak
sequence similarity that are not detectable during a comparison of
sequence. The files for this example are in the directory examples/s_prot in the
directory where you have installed STAMP.
A list of the domains is given in the file s_prot.domains. The
output. Note that you can create such a file by using the PDBC program.
Running PDBSEQ:
pdbseq -f s_prot.domains > s_prot.seqs
produced the file s_prot.seqs, from which an AMPS multiple
sequence alignment was produced, and stored in the file
s_prot_amps.align. Running ALIGNFIT:
alignfit -f s_prot_amps.align -d s_prot.domains -out s_prot_alignfit.trans
should give the output:
ALIGNFIT R.B. Russell 1995
Reading in block file...
Blocfile read: Length: 261
Reading in coordinate descriptions...
Reading coordinates...
Checking for inconsistencies...
Doing pairwise comparisons...
Doing treewise comparisons...
ALIGNFIT done.
Look in the file s_prot_alignfit.trans for output and details
The final transformation
(called alignfit.trans if default ALIGNFIT settings are used) is
in the file s_prot_alignfit.trans.
This provides an initial set of transformations for use by
STAMP. To run STAMP type:
stamp -l s_prot_alignfit.trans -prefix s_prot
Should produce the following output:
STAMP Structural Alignment of Multiple Proteins
by Robert B. Russell & Geoffrey J. Barton
Please cite PROTEINS, v14, 309-323, 1992
Sc = STAMP score, RMS = RMS deviation, Align = alignment length
Len1, Len2 = length of domain, Nfit = residues fitted
Secs = no. equivalent sec. strucs. Eq = no. equivalent residues
%I = seq. identity, %S = sec. str. identity
P(m) = P value (p=1/10) calculated after Murzin (1993), JMB, 230, 689-694
No. Domain1 Domain2 Sc RMS Len1 Len2 Align NFit Eq. Secs. %I %S P(m)
Pair 1 4chaa 3est 7.73 10.58 239 240 242 209 207 20 37.08 74.17 0.00e+00
Pair 2 4chaa 2ptn 7.80 9.43 239 223 234 201 198 20 40.17 75.31 0.00e+00
Pair 3 4chaa 1ton 6.81 9.38 239 227 245 183 178 19 29.71 66.11 2.19e-42
Pair 4 4chaa 3rp2a 7.45 9.95 239 224 235 195 188 18 29.71 67.78 2.36e-63
Pair 5 4chaa 2pkaab 7.27 9.65 239 232 241 198 195 19 29.71 73.64 0.00e+00
Pair 6 4chaa 1sgt 7.09 9.76 239 223 239 191 183 20 27.62 69.04 3.68e-49
Pair 7 4chaa 2sga 3.66 9.75 239 181 238 105 99 14 11.72 33.05 2.03e-07
Pair 8 4chaa 3sgbe 3.56 9.24 239 185 239 103 98 16 10.04 33.47 1.89e-05
Pair 9 4chaa 2alp 3.49 9.13 239 198 246 102 97 14 9.21 31.38 1.28e-04
<etc.>
Reading in matrix file s_prot.mat...
Doing cluster analysis...
Cluster: 1 ( 2ptn & 2pkaab ) Sc 8.50 RMS 10.20 Len 232 nfit 213
See file s_prot.1 for the alignment and transformations
Cluster: 2 ( 2sga & 3sgbe ) Sc 8.34 RMS 10.37 Len 191 nfit 164
See file s_prot.2 for the alignment and transformations
<etc.>
Cluster: 7 ( 2alp & 2sga 3sgbe ) Sc 8.40 RMS 10.30 Len 202 nfit 161
See file s_prot.7 for the alignment and transformations
Cluster: 8 ( 1sgt & 4chaa 3est 3rp2a 1ton 2ptn 2pkaab ) Sc 7.59 RMS 9.50 Len 268 nfit 175
See file s_prot.8 for the alignment and transformations
Cluster: 9 ( 1sgt 4chaa 3est 3rp2a 1ton 2ptn 2pkaab & 2alp 2sga 3sgbe ) Sc 4.77 RMS 9.74 Len 290 nfit 111
See file s_prot.9 for the alignment and transformations
The various fields describe details of the pairwise and treewise
comparisons: Sc, RMS deviation, the alignment length (Align),
the length of each structure in residues (Len1, Len2), the number of
atoms used in the RMS fit (Nfit), the number of equivalent secondary
structure elements (Secs), and the number of equivalent residues
(see above, Eq.).
STAMP will also produce several files:
s_prot.mat - a file containing the information used to derive the
structural similarity tree (i.e. the output from the PAIRWISE) mode. This
is an upper diagonal matrix containing the pairwise Sc values.
s_prot.N - a series of files containing transformations and alignments
created by running the TREEWISE mode in STAMP. Each file corresponds to
a node in the similarity tree (i.e. a cluster), where two
groups of one or more structures have been combined to form an
alignment and transformations. The higher the value of N the more
structurally dissimilar the proteins contained in the file are. Highly
similar structures are clustered (aligned/superimposed) at an early stage
in the program, with more distantly related structures being clustered
towards the end.
The top of each file contains the information needed to generate (using
TRANSFORM, see below)
superimposed coordinates (in STAMP domain format, see below).
After these details, various details of the similarity (RMS deviation,
Sc value, etc) are given. The bottom portion of the file contains
the structural alignment in STAMP format. Positions not aligned with
gaps contain information as to the degree of local structural similarity,
such as the distance between (averaged)
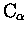 atoms, and the
atoms, and the
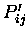 value.
value.
Methods for displaying sequence alignments and structures are described below.




Next: Alignment using an initial
Up: Worked examples
Previous: Worked examples
Geoff Barton
1999-04-16