


Next: VER2HOR
Up: Summary of parameters for
Previous: PDBSEQ
Contents
ALIGNFIT takes a multiple sequence alignment of proteins of known 3D structures
and uses it to superimpose them. It requires two files: an
AMPS multiple sequence alignment (block format), and a domain
description file. An optional parameter file may be supplied; if
none is given the program simply uses default parameters.
The format is:
alignfit -a <AMPS file> -d <domain file>
(-P <optional parm file> -<parameter> <value>)
-P can be used to read in an old ALIGNFIT parameter file (version 3.0 and earlier)
The possible parameters, and their defaults are (names are case
insensitive):
PAIRWISE  boolean
boolean
If TRUE, then pairwise comparisons will be performed to derive a
matrix (MATFILE).
Default PAIRWISE = TRUE
TREEWISE  boolean
boolean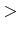
If TRUE, then treewise comparisons will be performed to derive a
final transformation.
Default TREEWISE = TRUE
MATFILE 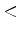 string
string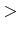
The file into which the results of PAIRWISE are output.
Default MATFILE = alignfit.mat
MAX_SEQ_LEN 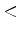 integer
integer
The maximum length of alignment to be tolerated.
Default is 3000
For most purposes, the default parameters should suffice. Note that one can
use ACONVERT to convert CLUSTAL and MSF formats to block format, so that one can use alignments created
using other programs (e.g. PILEUP, CLUSTAL, etc.) as a starting point for superimposition.



Next: VER2HOR
Up: Summary of parameters for
Previous: PDBSEQ
Contents