
Click on the protein name for more information about the protein.
The Score column is the predicted score of interaction between the two proteins
Click on the Evidence link to view a breakdown of the information about the predicted interaction.
| Key | |
|---|---|
| Module Score | Databases |
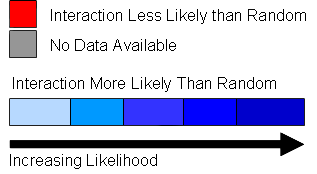 | Where interaction is observed in another database a link is provided: |
| Protein 1 (?) | Protein 2 (?) | Name of Protein 2 | Module Scores (?) | Scores (?) | Evidence (?) | Database (?) | |||
|---|---|---|---|---|---|---|---|---|---|
 |  |  |  | ||||||
| PSMD4 | MAPK1 | MAPK1, ERK2, PRKM1, PRKM2: Mitogen-activated protein kinase 1 | 496.0 | Evidence | |||||
| PSMD4 | NULL | N/A | 418.0 | Evidence | |||||
| PSMD4 | PSMD2 | PSMD2, TRAP2: 26S proteasome non-ATPase regulatory subunit 2 | 152.0 | Evidence | |||||
| PSMD4 | RAD23B | RAD23B: UV excision repair protein RAD23 homolog B | 86.80 | Evidence |

|
||||
| PSMD4 | PSMD11 | PSMD11: 26S proteasome non-ATPase regulatory subunit 11 | 52.00 | Evidence | |||||
| PSMD4 | RAD23A | RAD23A: UV excision repair protein RAD23 homolog A | 42.30 | Evidence |

|
||||
| PSMD4 | IQGAP1 | IQGAP1, KIAA0051: Ras GTPase-activating-like protein IQGAP1 | 25.50 | Evidence | |||||
| PSMD4 | SFRS10 | SFRS10, TRA2B: Splicing factor, arginine/serine-rich 10 | 25.00 | Evidence | |||||
| PSMD4 | COPS3 | COPS3, CSN3: COP9 signalosome complex subunit 3 | 20.60 | Evidence | |||||
| PSMD4 | TDG | TDG: G/T mismatch-specific thymine DNA glycosylase | 17.90 | Evidence | |||||
| PSMD4 | PSMC1 | PSMC1: 26S protease regulatory subunit 4 | 15.10 | Evidence | |||||
| PSMD4 | PSMC2 | PSMC2, MSS1: 26S protease regulatory subunit 7 | 15.10 | Evidence | |||||
| PSMD4 | USP14 | USP14, TGT: Ubiquitin carboxyl-terminal hydrolase 14 | 13.40 | Evidence | |||||
| PSMD4 | IQGAP2 | IQGAP2: Ras GTPase-activating-like protein IQGAP2 | 13.20 | Evidence | |||||