
Click on the protein name for more information about the protein.
The Score column is the predicted score of interaction between the two proteins
Click on the Evidence link to view a breakdown of the information about the predicted interaction.
| Key | |
|---|---|
| Module Score | Databases |
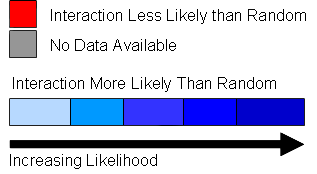 | Where interaction is observed in another database a link is provided: |
| Protein 1 (?) | Protein 2 (?) | Name of Protein 2 | Module Scores (?) | Scores (?) | Evidence (?) | Database (?) | |||
|---|---|---|---|---|---|---|---|---|---|
 |  |  |  | ||||||
| BTK | JAK2 | JAK2: Tyrosine-protein kinase JAK2 | 835.0 | Evidence | |||||
| BTK | TYK2 | TYK2: Non-receptor tyrosine-protein kinase TYK2 | 720.0 | Evidence | |||||
| BTK | SYK | SYK: Tyrosine-protein kinase SYK | 720.0 | Evidence |

|
||||
| BTK | CRKL | CRKL: Crk-like protein | 674.0 | Evidence | |||||
| BTK | LYN | LYN: Tyrosine-protein kinase Lyn | 613.0 | Evidence |

|
||||
| BTK | FGR | FGR, SRC2: Proto-oncogene tyrosine-protein kinase FGR | 565.0 | Evidence | |||||
| BTK | CSK | CSK: Tyrosine-protein kinase CSK | 549.0 | Evidence | |||||
| BTK | STAT5A | STAT5A, STAT5: Signal transducer and activator of transcription 5A | 497.0 | Evidence |

|
||||
| BTK | HCK | N/A | 392.0 | Evidence |
 O
O
|
||||
| BTK | BLNK | BLNK, BASH, SLP65: B-cell linker protein | 370.0 | Evidence |

|
||||
| BTK | VAV1 | VAV1, VAV: Proto-oncogene vav | 332.0 | Evidence |
 O
O
|
||||
| BTK | STAT5B | STAT5B: Signal transducer and activator of transcription 5B | 305.0 | Evidence | |||||
| BTK | SH3BP2 | SH3BP2: Uncharacterized protein SH3BP2 | 299.0 | Evidence | |||||
| BTK | FYN | FYN: Proto-oncogene tyrosine-protein kinase Fyn | 299.0 | Evidence | |||||
| BTK | NCK1 | NCK1, NCK: Cytoplasmic protein NCK1 | 259.0 | Evidence | |||||
| BTK | KIT | KIT: Mast/stem cell growth factor receptor precursor | 206.0 | Evidence |

|
||||
| BTK | APS | SH2B2, APS: SH2B adapter protein 2 | 198.0 | Evidence |

|
||||
| BTK | SRC | SRC, SRC1: Proto-oncogene tyrosine-protein kinase Src | 181.0 | Evidence | |||||
| BTK | SLA | SLA, SLAP, SLAP1: SRC-like-adapter | 181.0 | Evidence | |||||
| BTK | LCP2 | LCP2: Lymphocyte cytosolic protein 2 | 177.0 | Evidence | |||||
| BTK | JAK1 | 162.0 | Evidence |
 O
O
|
|||||
| BTK | STAT3 | 156.0 | Evidence | ||||||
| BTK | PDGFRB | PDGFRB: Beta-type platelet-derived growth factor receptor precursor | 149.0 | Evidence | |||||
| BTK | GRB2 | GRB2, ASH: Growth factor receptor-bound protein 2 | 139.0 | Evidence | O | ||||
| BTK | STAT1 | STAT1: Signal transducer and activator of transcription 1-alpha/beta | 135.0 | Evidence | |||||
| BTK | CSF1R | CSF1R, FMS: Macrophage colony-stimulating factor 1 receptor precursor | 135.0 | Evidence | |||||
| BTK | ZAP70 | ZAP70: Uncharacterized protein ZAP70 | 121.0 | Evidence | |||||
| BTK | ABL2 | ABL2, ABLL, ARG: Tyrosine-protein kinase ABL2 | 119.0 | Evidence | |||||
| BTK | PTPN6 | PTPN6, HCP, PTP1C: Tyrosine-protein phosphatase non-receptor type 6 | 119.0 | Evidence | |||||
| BTK | ABL1 | ABL1, ABL, JTK7: Proto-oncogene tyrosine-protein kinase ABL1 | 109.0 | Evidence |

|
||||
| BTK | PTK2B | PTK2B, FAK2, PYK2, RAFTK: Protein tyrosine kinase 2 beta | 106.0 | Evidence | |||||
| BTK | RASA1 | RASA1, RASA: Ras GTPase-activating protein 1 | 106.0 | Evidence | |||||
| BTK | ITK | ITK, EMT, LYK: Tyrosine-protein kinase ITK/TSK | 106.0 | Evidence |

|
||||
| BTK | TEC | TEC, PSCTK4: Tyrosine-protein kinase Tec | 98.20 | Evidence |

|
||||
| BTK | WASPIP | N/A | 95.20 | Evidence | |||||
| BTK | CBLB | CBLB, RNF56, Nbla00127: E3 ubiquitin-protein ligase CBL-B | 92.20 | Evidence | |||||
| BTK | PDGFRA | PDGFRA: Alpha-type platelet-derived growth factor receptor precursor | 90.70 | Evidence | |||||
| BTK | LCK | LCK: Proto-oncogene tyrosine-protein kinase LCK | 88.80 | Evidence | |||||
| BTK | KHDRBS1 | KHDRBS1, SAM68: KH domain-containing, RNA-binding, signal transduction-associate | 87.80 | Evidence |

|
||||
| BTK | KDR | KDR, FLK1: Vascular endothelial growth factor receptor 2 precursor | 80.00 | Evidence | |||||
| BTK | INSR | INSR: Insulin receptor precursor | 80.00 | Evidence | |||||
| BTK | INSL3 | JAK3: Tyrosine-protein kinase JAK3 | 78.20 | Evidence | |||||
| BTK | PTK6 | PTK6, BRK: Tyrosine-protein kinase 6 | 78.20 | Evidence | |||||
| BTK | BMX | N/A | 78.20 | Evidence |

|
||||
| BTK | CBL | CBL, CBL2, RNF55: E3 ubiquitin-protein ligase CBL | 78.20 | Evidence |
 O
O
|
||||
| BTK | MAP4K1 | MAP4K1, HPK1: Mitogen-activated protein kinase kinase kinase kinase 1 | 77.50 | Evidence | |||||
| BTK | PIK3R2 | PIK3R2: Phosphatidylinositol 3-kinase regulatory subunit beta | 70.10 | Evidence | |||||
| BTK | HCLS1 | HCLS1, HS1: Hematopoietic lineage cell-specific protein | 69.40 | Evidence | |||||
| BTK | TEK | TEK, TIE2: Angiopoietin-1 receptor precursor | 59.50 | Evidence | |||||
| BTK | WAS | WAS, IMD2: Wiskott-Aldrich syndrome protein | 55.10 | Evidence |
 O
O
|
||||
| BTK | CBLC | CBLC, CBL3, RNF57: Signal transduction protein CBL-C | 52.80 | Evidence | |||||
| BTK | EPHB3 | EPHB3, ETK2, HEK2, TYRO6: Ephrin type-B receptor 3 precursor | 47.30 | Evidence | |||||
| BTK | CDK7 | CDK7, MO15: Cell division protein kinase 7 | 38.10 | Evidence | |||||
| BTK | PIK3CG | PIK3CG: Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma i | 37.80 | Evidence | |||||
| BTK | GRAP | GRAP: GRB2-related adapter protein | 36.40 | Evidence | |||||
| BTK | NMT1 | NMT1, NMT: Glycylpeptide N-tetradecanoyltransferase 1 | 35.00 | Evidence | |||||
| BTK | DOK1 | DOK1: Docking protein 1 | 33.90 | Evidence | |||||
| BTK | SOCS1 | SOCS1, SSI1, TIP3: Suppressor of cytokine signaling 1 | 32.80 | Evidence | |||||
| BTK | CDK2 | CDK2: Cell division protein kinase 2 | 25.00 | Evidence | |||||
| BTK | CD22 | CD22, SIGLEC2: B-cell receptor CD22 precursor | 24.90 | Evidence | |||||
| BTK | CD79A | CD79A, IGA, MB1: B-cell antigen receptor complex-associated protein alpha-chain | 24.90 | Evidence | |||||
| BTK | TUBA1 | TUBA4A, TUBA1: Tubulin alpha-4A chain | 24.90 | Evidence | |||||
| BTK | BRDG1 | STAP1, BRDG1: Signal-transducing adaptor protein 1 | 24.10 | Evidence | |||||
| BTK | ACTG1 | ACTB: Actin, cytoplasmic 1 | 21.20 | Evidence | |||||
| BTK | NRBP1 | NRBP1, BCON3, NRBP: Nuclear receptor-binding protein | 20.50 | Evidence | |||||
| BTK | PRKDC | PRKDC, HYRC, HYRC1: DNA-dependent protein kinase catalytic subunit | 19.10 | Evidence | |||||
| BTK | CXCR4 | CXCR4: C-X-C chemokine receptor type 4 | 18.40 | Evidence | |||||
| BTK | IL10RA | IL10RA, IL10R: Interleukin-10 receptor alpha chain precursor | 18.40 | Evidence | |||||
| BTK | CSF2RB | CSF2RB, IL3RB, IL5RB: Cytokine receptor common beta chain precursor | 18.40 | Evidence | |||||
| BTK | SOCS5 | SOCS5, CIS6, CISH5, CISH6, KIAA0671: Suppressor of cytokine signaling 5 | 15.80 | Evidence | |||||
| BTK | SNCA | SNCA, NACP, PARK1: Alpha-synuclein | 15.00 | Evidence | |||||
| BTK | CDC2 | CDC2: Cell division control protein 2 homolog | 15.00 | Evidence | |||||
| BTK | DUSP16 | DUSP16, KIAA1700, MKP7: Dual specificity protein phosphatase 16 | 15.00 | Evidence | |||||
| BTK | SH3KBP1 | SH3KBP1, CIN85: SH3 domain-containing kinase-binding protein 1 | 15.00 | Evidence | |||||
| BTK | PASK | PASK, KIAA0135: PAS domain-containing serine/threonine-protein kinase | 13.80 | Evidence | |||||
| BTK | PTPN2 | PTPN2, PTPT: Tyrosine-protein phosphatase non-receptor type 2 | 13.50 | Evidence | |||||
| BTK | SOS1 | SOS1: Son of sevenless homolog 1 | 13.50 | Evidence | |||||
| BTK | NEDD9 | NEDD9, CASL: Enhancer of filamentation 1 | 13.40 | Evidence | |||||
| BTK | MAPK3 | MAPK3, ERK1, PRKM3: Mitogen-activated protein kinase 3 | 13.2 | Evidence | |||||
| BTK | AMHR2 | AMHR2, AMHR: Anti-Muellerian hormone type-2 receptor precursor | 13.00 | Evidence | |||||
| BTK | SCAP2 | SKAP2, PRAP, RA70, SAPS, SCAP2, SKAP55R: Src kinase-associated phosphoprotein 2 | 12.50 | Evidence | |||||
| BTK | EPOR | EPOR: Erythropoietin receptor precursor | 11.90 | Evidence | |||||
| BTK | PECAM1 | PECAM1: Platelet endothelial cell adhesion molecule precursor | 11.70 | Evidence | |||||
| BTK | QARS | QARS: Glutaminyl-tRNA synthetase | 11.70 | Evidence | |||||
| BTK | HNRPK | N/A | 11.70 | Evidence | |||||
| BTK | ICOSLG | ICOSLG, B7H2, B7RP1, ICOSL, KIAA0653: ICOS ligand precursor | 11.40 | Evidence | |||||
| BTK | PRKAA1 | N/A | 11.20 | Evidence | |||||
| BTK | PTPRC | PTPRC, RP11-553K8.4-001, hCG_1811178: Protein tyrosine phosphatase, receptor typ | 9.92 | Evidence | |||||
| BTK | IL6ST | IL6ST: Interleukin-6 receptor subunit beta precursor | 9.92 | Evidence | |||||
| BTK | ADAM15 | ADAM15, MDC15: ADAM 15 precursor | 9.92 | Evidence | |||||
| BTK | LAT | LAT: Linker for activation of T-cells family member 1 | 9.92 | Evidence | |||||
| BTK | CD79B | CD79B, B29, IGB: B-cell antigen receptor complex-associated protein beta-chain p | 9.92 | Evidence | |||||
| BTK | TRPV4 | TRPV4, VRL2, VROAC: Transient receptor potential cation channel subfamily V memb | 9.92 | Evidence | |||||
| BTK | FCGR2B | FCGR2B, CD32, FCG2, IGFR2: Low affinity immunoglobulin gamma Fc region receptor | 9.92 | Evidence | |||||
| BTK | PAK7 | PAK7, KIAA1264, PAK5: Serine/threonine-protein kinase PAK 7 | 9.0 | Evidence | |||||
| BTK | PRKD1 | PRKD1, PKD, PKD1, PRKCM: Serine/threonine-protein kinase D1 | 9.0 | Evidence |
 O
O
|
||||
| BTK | MYL9 | MYL9, MLC2, MRLC1, MYRL2: Myosin regulatory light chain 2, smooth muscle isoform | 9.0 | Evidence | |||||
| BTK | TRPM7 | TRPM7, CHAK1, LTRPC7: Transient receptor potential cation channel subfamily M me | 8.89 | Evidence | |||||
| BTK | CAMK1 | CAMK1: Calcium/calmodulin-dependent protein kinase type 1 | 8.86 | Evidence | |||||
| BTK | PRKCA | PRKCA, PKCA, PRKACA: Protein kinase C alpha type | 8.86 | Evidence |

|
||||
| BTK | ITGB1BP1 | ITGB1BP1, ICAP1: Integrin beta-1-binding protein 1 | 8.84 | Evidence | |||||
| BTK | GNA15 | GNA15, GNA16: Guanine nucleotide-binding protein alpha-15 subunit | 8.31 | Evidence | |||||
| BTK | PRKCQ | PRKCQ, PRKCT: Protein kinase C theta type | 7.99 | Evidence |

|
||||
| BTK | CTTN | CTTN, EMS1: Src substrate cortactin | 7.97 | Evidence | |||||
| BTK | DAPP1 | N/A | 7.93 | Evidence |

|
||||
| BTK | UBAP2L | UBAP2L, KIAA0144, NICE4: Ubiquitin-associated protein 2-like | 7.93 | Evidence | |||||
| BTK | VAV2 | VAV2, RP11-153P4.1-003: Vav 2 oncogene | 7.93 | Evidence | |||||
| BTK | EIF4EBP1 | EIF4EBP1: Eukaryotic translation initiation factor 4E-binding protein 1 | 7.79 | Evidence | |||||
| BTK | HNRPK | HNRPK, HNRNPK: Heterogeneous nuclear ribonucleoprotein K | 7.59 | Evidence | |||||
| BTK | CASP9 | CASP9, MCH6: Caspase-9 precursor | 7.59 | Evidence | |||||
| BTK | IL2RB | IL2RB: Interleukin-2 receptor subunit beta precursor | 7.40 | Evidence | |||||
| BTK | STAG2 | STAG2, SA2: Cohesin subunit SA-2 | 7.10 | Evidence | |||||
| BTK | CD36 | CD36, GP3B, GP4: Platelet glycoprotein 4 | 6.81 | Evidence | |||||
| BTK | GIT2 | N/A | 6.80 | Evidence | |||||
| BTK | CD3E | CD3E, T3E: T-cell surface glycoprotein CD3 epsilon chain precursor | 6.52 | Evidence | |||||
| BTK | IL1RL1 | IL1RL1, DER4, ST2, T1: Interleukin-1 receptor-like 1 precursor | 6.52 | Evidence | |||||
| BTK | GNA12 | GNA12: Guanine nucleotide-binding protein alpha-12 subunit | 6.1 | Evidence |

|
||||
| BTK | ARPC1B | ARC41, ARPC1B: Actin-related protein 2/3 complex subunit 1B | 6.01 | Evidence | |||||
| BTK | DUSP3 | DUSP3, VHR: Dual specificity protein phosphatase 3 | 5.88 | Evidence | |||||
| BTK | ITGB4 | N/A | 5.88 | Evidence | |||||
| BTK | MIP | MIP, AQP0: Lens fiber major intrinsic protein | 5.84 | Evidence | |||||
| BTK | HSPD1 | 5.74 | Evidence | ||||||
| BTK | PPP1R12A | PPP1R12A, MBS, MYPT1: Protein phosphatase 1 regulatory subunit 12A | 5.74 | Evidence | |||||
| BTK | STAT6 | STAT6: Signal transducer and activator of transcription 6 | 5.22 | Evidence | |||||
| BTK | HLA-F | HLA-F, HLA-5.4, HLAF: HLA class I histocompatibility antigen, alpha chain F prec | 4.80 | Evidence | |||||
| BTK | IFNAR2 | N/A | 4.80 | Evidence | |||||
| BTK | FCER1G | FCER1G: High affinity immunoglobulin epsilon receptor subunit gamma precursor | 4.80 | Evidence | |||||
| BTK | DNM2 | N/A | 4.73 | Evidence | |||||
| BTK | IGHE | IGHE: Ig epsilon chain C region | 4.68 | Evidence | |||||
| BTK | CDC42BPA | CDC42BPA: Serine/threonine-protein kinase MRCK alpha | 4.68 | Evidence | |||||
| BTK | PLAUR | PLAUR, MO3, UPAR: Urokinase plasminogen activator surface receptor precursor | 4.48 | Evidence | |||||
| BTK | TNKS2 | TNKS2, PARP5B, TANK2, TNKL: Tankyrase-2 | 4.30 | Evidence | |||||
| BTK | BCAR1 | BCAR1, CAS, CRKAS: Breast cancer anti-estrogen resistance protein 1 | 4.19 | Evidence | |||||
| BTK | CD19 | N/A | 4.06 | Evidence |

|
||||
| BTK | PRKX | PRKX, PKX1: Serine/threonine-protein kinase PRKX | 4.04 | Evidence | |||||
| BTK | MTMR1 | MTMR1: Myotubularin-related protein 1 | 4.04 | Evidence | |||||
| BTK | STOM | STOM, BND7, EPB72: Erythrocyte band 7 integral membrane protein | 4.01 | Evidence | |||||
| BTK | GRIA3 | GRIA3, GLUR3, GLURC: Glutamate receptor 3 precursor | 4.01 | Evidence | |||||
| BTK | GTF2B | GTF2B, TF2B, TFIIB: Transcription initiation factor IIB | 3.64 | Evidence | |||||
| BTK | PPP1R8 | N/A | 3.64 | Evidence | |||||
| BTK | FCGR1A | FCGR1A, FCG1, FCGR1, IGFR1: High affinity immunoglobulin gamma Fc receptor I pre | 3.54 | Evidence | |||||
| BTK | CASP3 | CASP3, CPP32: Caspase-3 precursor | 3.47 | Evidence | |||||
| BTK | LYPD3 | LYPD3, C4.4A, UNQ491/PRO1007: Ly6/PLAUR domain-containing protein 3 precursor | 2.94 | Evidence | |||||
| BTK | ZYX | ZYX: Zyxin | 2.94 | Evidence | |||||
| BTK | SLC9A3R1 | SLC9A3R1, NHERF: Ezrin-radixin-moesin-binding phosphoprotein 50 | 2.94 | Evidence | |||||
| BTK | DHX16 | DHX16, DBP2, DDX16, KIAA0577: Putative pre-mRNA-splicing factor ATP-dependent RN | 2.93 | Evidence | |||||
| BTK | BFAR | BFAR, BAR, RNF47: Bifunctional apoptosis regulator | 2.90 | Evidence | |||||
| BTK | AQP2 | AQP2: Aquaporin-2 | 2.80 | Evidence | |||||
| BTK | CTLA4 | CTLA4, CD152: Cytotoxic T-lymphocyte protein 4 precursor | 2.77 | Evidence | |||||
| BTK | AGTR1 | AGTR1, AGTR1A, AGTR1B, AT2R1, AT2R1B: Type-1 angiotensin II receptor | 2.65 | Evidence | |||||
| BTK | TSHR | N/A | 2.63 | Evidence | |||||
| BTK | KCNJ12 | KCNJ12, IRK2, KCNJN1: ATP-sensitive inward rectifier potassium channel 12 | 2.63 | Evidence | |||||
| BTK | TXN | TXN, TRDX, TRX, TRX1: Thioredoxin | 2.63 | Evidence | |||||
| BTK | MATK | 2.54 | Evidence | ||||||
| BTK | NASP | NASP: Nuclear autoantigenic sperm protein | 2.54 | Evidence | |||||