
Click on the protein name for more information about the protein.
The Score column is the predicted score of interaction between the two proteins
Click on the Evidence link to view a breakdown of the information about the predicted interaction.
| Key | |
|---|---|
| Module Score | Databases |
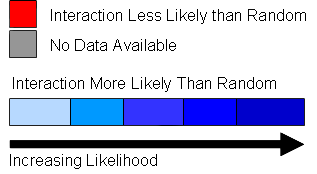 | Where interaction is observed in another database a link is provided: |
| Protein 1 (?) | Protein 2 (?) | Name of Protein 2 | Module Scores (?) | Scores (?) | Evidence (?) | Database (?) | |||
|---|---|---|---|---|---|---|---|---|---|
 |  |  |  | ||||||
| MCM2 | MCM4 | MCM4, CDC21: DNA replication licensing factor MCM4 | 2.96E5 | Evidence | |||||
| MCM2 | MCM3 | MCM3: DNA replication licensing factor MCM3 | 2.96E5 | Evidence |

 O
O
|
||||
| MCM2 | CDC6 | CDC6, CDC18L: Cell division control protein 6 homolog | 9.88E4 | Evidence |

 O
O
|
||||
| MCM2 | MCM7 | MCM7, CDC47, MCM2: DNA replication licensing factor MCM7 | 4.78E4 | Evidence |


|
||||
| MCM2 | MCM5 | MCM5, CDC46: DNA replication licensing factor MCM5 | 4.78E4 | Evidence |

 O
O
|
||||
| MCM2 | RPA1 | RPA1, REPA1, RPA70: Replication protein A 70 kDa DNA-binding subunit | 1.59E4 | Evidence |
 O
O
|
||||
| MCM2 | MCM6 | MCM6: DNA replication licensing factor MCM6 | 1.59E4 | Evidence |

 O
O
|
||||
| MCM2 | CDC7 | CDC7, CDC7L1: Cell division cycle 7-related protein kinase | 1.46E4 | Evidence |

|
||||
| MCM2 | CCNB1 | Cyclin B1 | 372.0 | Evidence | |||||
| MCM2 | ORC5L | ORC5L: Origin recognition complex subunit 5 | 341.0 | Evidence |

 O
O
|
||||
| MCM2 | PCNA | PCNA: Proliferating cell nuclear antigen | 189.0 | Evidence | |||||
| MCM2 | ORC1L | ORC1L, ORC1, PARC1: Origin recognition complex subunit 1 | 174.0 | Evidence |
 O
O
|
||||
| MCM2 | CDK2 | CDK2: Cell division protein kinase 2 | 117.0 | Evidence | |||||
| MCM2 | RB1 | RB1: Retinoblastoma-associated protein | 117.0 | Evidence | |||||
| MCM2 | BRCA2 | BRCA2, FACD, FANCD1: Breast cancer type 2 susceptibility protein | 117.0 | Evidence | |||||
| MCM2 | WRN | WRN, RECQ3, RECQL2: Werner syndrome ATP-dependent helicase | 105.0 | Evidence | |||||
| MCM2 | BLM | BLM, RECQ2, RECQL3: Bloom syndrome protein | 105.0 | Evidence | |||||
| MCM2 | CCNF | CCNF: G2/mitotic-specific cyclin-F | 67.20 | Evidence | |||||
| MCM2 | CSDA | CSDA, DBPA: DNA-binding protein A | 62.90 | Evidence | |||||
| MCM2 | CDK4 | CDK4: Cell division protein kinase 4 | 61.50 | Evidence |

 O
O
|
||||
| MCM2 | CCNA2 | CCNA2, CCN1, CCNA: Cyclin-A2 | 60.20 | Evidence | |||||
| MCM2 | GTF2B | GTF2B, TF2B, TFIIB: Transcription initiation factor IIB | 51.60 | Evidence |

|
||||
| MCM2 | RPA2 | RPA2, REPA2, RPA32, RPA34: Replication protein A 32 kDa subunit | 51.50 | Evidence |

|
||||
| MCM2 | PLK4 | PLK4, SAK, STK18: Serine/threonine-protein kinase PLK4 | 47.30 | Evidence | |||||
| MCM2 | CDC45L | CDC45L, CDC45L2, UNQ374/PRO710: CDC45-related protein | 44.90 | Evidence | |||||
| MCM2 | BIRC5 | BIRC5, API4, IAP4: Baculoviral IAP repeat-containing protein 5 | 36.00 | Evidence | |||||
| MCM2 | VCP | VCP: Transitional endoplasmic reticulum ATPase | 34.00 | Evidence | |||||
| MCM2 | PLK1 | PLK1, PLK: Serine/threonine-protein kinase PLK1 | 33.90 | Evidence |

|
||||
| MCM2 | SNF1LK2 | SNF1LK2, KIAA0781, QIK: Serine/threonine-protein kinase SNF1-like kinase 2 | 33.40 | Evidence | |||||
| MCM2 | HDAC3 | HDAC3: Histone deacetylase 3 | 29.50 | Evidence | |||||
| MCM2 | NPM1 | NPM1, NPM: Nucleophosmin | 29.50 | Evidence | |||||
| MCM2 | TP53 | TP53, P53: Cellular tumor antigen p53 | 28.80 | Evidence | |||||
| MCM2 | ORC2L | ORC2L, ORC2: Origin recognition complex subunit 2 | 28.2 | Evidence |

 O
O
|
||||
| MCM2 | ORC6L | ORC6L: Origin recognition complex subunit 6 | 28.2 | Evidence |

 O
O
|
||||
| MCM2 | TFDP1 | TFDP1, DP1: Transcription factor Dp-1 | 28.2 | Evidence | |||||
| MCM2 | SFRS6 | SFRS6, SRP55: Splicing factor, arginine/serine-rich 6 | 26.90 | Evidence | |||||
| MCM2 | POLR2A | POLR2A: Uncharacterized protein POLR2A | 26.00 | Evidence | |||||
| MCM2 | SMARCA4 | SMARCA4, BRG1, SNF2B, SNF2L4: Probable global transcription activator SNF2L4 | 25.00 | Evidence | |||||
| MCM2 | CCNH | CCNH: Cyclin-H | 23.70 | Evidence | |||||
| MCM2 | HMGN1 | HMGN1, HMG14: Non-histone chromosomal protein HMG-14 | 19.90 | Evidence | |||||
| MCM2 | PABPC1 | PABPC1, PAB1, PABP1, PABPC2: Polyadenylate-binding protein 1 | 19.90 | Evidence | |||||
| MCM2 | RPS6KB1 | N/A | 19.90 | Evidence | |||||
| MCM2 | HDAC2 | HDAC2: Histone deacetylase 2 | 17.80 | Evidence | |||||
| MCM2 | RAD51 | RAD51, RAD51A, RECA: DNA repair protein RAD51 homolog 1 | 17.60 | Evidence | |||||
| MCM2 | TCOF1 | N/A | 16.10 | Evidence | |||||
| MCM2 | CCND1 | CCND1, BCL1, PRAD1: G1/S-specific cyclin-D1 | 15.30 | Evidence | |||||
| MCM2 | AKAP8L | AKAP8L, NAKAP95, HRIHFB2018: A-kinase anchor protein 8-like | 14.40 | Evidence | |||||
| MCM2 | SCYE1 | SCYE1, EMAP2: Multisynthetase complex auxiliary component p43 [Contains: Endothe | 13.70 | Evidence | |||||
| MCM2 | ARFGAP1 | N/A | 11.80 | Evidence | |||||
| MCM2 | MUTYH | N/A | 11.50 | Evidence | |||||
| MCM2 | STAT1 | STAT1: Signal transducer and activator of transcription 1-alpha/beta | 10.70 | Evidence | |||||
| MCM2 | DYRK1B | DYRK1B, MIRK: Dual specificity tyrosine-phosphorylation-regulated kinase 1B | 10.70 | Evidence | |||||
| MCM2 | TUBA1 | TUBA4A, TUBA1: Tubulin alpha-4A chain | 10.60 | Evidence | |||||
| MCM2 | SMAD2 | SMAD2, MADH2, MADR2: Mothers against decapentaplegic homolog 2 | 10.60 | Evidence | |||||
| MCM2 | CRSP2 | MED14, ARC150, CRSP2, CXorf4, DRIP150, EXLM1, RGR1, TRAP170: Mediator of RNA pol | 8.72 | Evidence | |||||
| MCM2 | PHB2 | PHB2, BAP, REA: Prohibitin-2 | 8.24 | Evidence | |||||
| MCM2 | GMNN | GMNN: Geminin | 7.42 | Evidence | |||||
| MCM2 | HIVEP1 | N/A | 7.39 | Evidence | |||||
| MCM2 | MTF1 | MTF1: Metal regulatory transcription factor 1 | 6.62 | Evidence | |||||
| MCM2 | AURKB | AURKB, AIK2, AIM1, ARK2, STK12: Serine/threonine-protein kinase 12 | 6.35 | Evidence | |||||
| MCM2 | KIAA0101 | PAF, KIAA0101, L5, NS5ATP9: PCNA-associated factor | 6.29 | Evidence | |||||
| MCM2 | RECQL | RECQL, RECQL1: ATP-dependent DNA helicase Q1 | 6.16 | Evidence | |||||
| MCM2 | TRAF2 | TRAF2, TRAP3: TNF receptor-associated factor 2 | 6.08 | Evidence | |||||
| MCM2 | GEMIN5 | GEMIN5: Gem-associated protein 5 | 5.73 | Evidence | |||||
| MCM2 | BCOR | N/A | 5.73 | Evidence | |||||
| MCM2 | CDC25A | CDC25A: M-phase inducer phosphatase 1 | 5.73 | Evidence | |||||
| MCM2 | PHC2 | PHC2, EDR2, PH2: Polyhomeotic-like protein 2 | 5.73 | Evidence |

|
||||
| MCM2 | NUP98 | N/A | 5.73 | Evidence | |||||
| MCM2 | TCEA1 | N/A | 5.73 | Evidence | |||||
| MCM2 | DBF4 | DBF4, ASK: Protein DBF4 homolog | 5.73 | Evidence |

|
||||
| MCM2 | HAT1 | HAT1: Histone acetyltransferase type B catalytic subunit | 5.64 | Evidence | O | ||||
| MCM2 | OBFC1 | OBFC1: Oligonucleotide/oligosaccharide-binding fold-containing protein 1 | 5.63 | Evidence | |||||
| MCM2 | CCND2 | CCND2: G1/S-specific cyclin-D2 | 5.30 | Evidence | |||||
| MCM2 | DIAPH1 | Diaphanous 1 | 5.28 | Evidence | |||||
| MCM2 | GIT1 | GIT1: ARF GTPase-activating protein GIT1 | 5.28 | Evidence | |||||
| MCM2 | DDX1 | DDX1: ATP-dependent RNA helicase DDX1 | 5.24 | Evidence | |||||
| MCM2 | DDX18 | DDX18: ATP-dependent RNA helicase DDX18 | 5.23 | Evidence | |||||
| MCM2 | CDT1 | CDT1: DNA replication factor Cdt1 | 5.13 | Evidence | |||||
| MCM2 | ID2 | ID2: DNA-binding protein inhibitor ID-2 | 4.77 | Evidence | |||||
| MCM2 | KLF13 | KLF13, BTEB3, NSLP1: Krueppel-like factor 13 | 4.55 | Evidence | |||||
| MCM2 | CTNNB1 | CTNNB1, CTNNB, OK/SW-cl.35, PRO2286: Catenin beta-1 | 4.46 | Evidence | |||||
| MCM2 | FYN | FYN: Proto-oncogene tyrosine-protein kinase Fyn | 4.28 | Evidence | |||||
| MCM2 | RPA3 | RPA3, REPA3, RPA14: Replication protein A 14 kDa subunit | 3.70 | Evidence |
 O
O
|
||||
| MCM2 | NEDD4 | NEDD4: Uncharacterized protein NEDD4 | 3.47 | Evidence | |||||
| MCM2 | ZNF174 | ZNF174, ZSCAN8: Zinc finger protein 174 | 3.47 | Evidence | |||||
| MCM2 | CAMK2A | N/A | 3.47 | Evidence | |||||
| MCM2 | ARID4A | ARID4A, RBBP1, RBP1: AT-rich interactive domain-containing protein 4A | 3.47 | Evidence | |||||
| MCM2 | CDKN1A | CDKN1A, CAP20, CDKN1, CIP1, MDA6, PIC1, SDI1, WAF1: Cyclin-dependent kinase inhi | 2.89 | Evidence | |||||
| MCM2 | HMGA2 | HMGA2, HMGIC: High mobility group protein HMGI-C | 2.89 | Evidence | |||||
| MCM2 | ABCD3 | ABCD3, PMP70, PXMP1: ATP-binding cassette sub-family D member 3 | 2.74 | Evidence | |||||
| MCM2 | CASP2 | CASP2, ICH1, NEDD2: Caspase-2 precursor | 2.70 | Evidence | |||||
| MCM2 | CREBBP | CREBBP, CBP: CREB-binding protein | 2.10 | Evidence | |||||
| MCM2 | RP2 | RP2: Protein XRP2 | 2.05 | Evidence | |||||
| MCM2 | SH2D3A | SH2D3A, NSP1, UNQ175/PRO201: SH2 domain-containing protein 3A | 1.98 | Evidence | |||||
| MCM2 | C10orf117 | NOC3L, AD24, C10orf117, FAD24: Nucleolar complex protein 3 homolog | 1.82 | Evidence | |||||
| MCM2 | RNF138 | N/A | 1.78 | Evidence | |||||
| MCM2 | WRNIP1 | WRNIP1, WHIP: ATPase WRNIP1 | 1.78 | Evidence | |||||
| MCM2 | ORC4L | ORC4L: Origin recognition complex subunit 4 | 1.71 | Evidence |

|
||||
| MCM2 | POLE | POLE, POLE1: DNA polymerase epsilon catalytic subunit A | 1.57 | Evidence | |||||
| MCM2 | LIAS | LIAS, LAS, HUSSY-01: Lipoic acid synthetase, mitochondrial precursor | 1.10 | Evidence | O | ||||
| MCM2 | TOP2A | TOP2A, TOP2: DNA topoisomerase 2-alpha | 0.797 | Evidence | |||||
| MCM2 | E2F1 | E2F1, RBBP3: Transcription factor E2F1 | 0.738 | Evidence | |||||
| MCM2 | HMGB2 | HMGB2, HMG2: High mobility group protein B2 | 0.715 | Evidence | |||||
| MCM2 | EIF2S1 | EIF2S1, EIF2A: Eukaryotic translation initiation factor 2 subunit 1 | 0.66 | Evidence | |||||
| MCM2 | ORC3L | ORC3L, LATHEO: Origin recognition complex subunit 3 | 0.632 | Evidence | |||||
| MCM2 | MCM10 | MCM10, PRO2249: Protein MCM10 homolog | 0.593 | Evidence |

|
||||
| MCM2 | MSH3 | MSH3, DUC1, DUG: DNA mismatch repair protein Msh3 | 0.472 | Evidence | |||||
| MCM2 | CHEK1 | CHEK1, CHK1: Serine/threonine-protein kinase Chk1 | 0.445 | Evidence | |||||
| MCM2 | VRK1 | VRK1: Serine/threonine-protein kinase VRK1 | 0.445 | Evidence | |||||
| MCM2 | AURKA | AURKA, AIK, ARK1, AURA, BTAK, STK15, STK6: Serine/threonine-protein kinase 6 | 0.445 | Evidence | |||||
| MCM2 | SFRS1 | SFRS1, ASF, OK/SW-cl.3, SF2, SF2P33: Splicing factor, arginine/serine-rich 1 | 0.445 | Evidence | |||||
| MCM2 | DNMT1 | DNMT1, AIM, DNMT: DNA | 0.445 | Evidence | |||||
| MCM2 | EFNB1 | EFNB1, EFL-3, EPLG2, LERK2: Ephrin-B1 precursor | 0.438 | Evidence | |||||
| MCM2 | ATAD2 | ATAD2, L16, PRO2000: ATPase family AAA domain-containing protein 2 | 0.435 | Evidence | |||||
| MCM2 | YBX1 | YBX1, NSEP1, YB1: Nuclease sensitive element-binding protein 1 | 0.4 | Evidence | |||||
| MCM2 | EIF2S2 | EIF2S2, EIF2B: Eukaryotic translation initiation factor 2 subunit 2 | 0.4 | Evidence | |||||
| MCM2 | GTF2F1 | GTF2F1, RAP74: Transcription initiation factor IIF subunit alpha | 0.4 | Evidence | |||||
| MCM2 | LHB | LHB: Lutropin subunit beta precursor | 0.395 | Evidence | |||||
| MCM2 | ADPRT | PARP1, ADPRT, PPOL: Poly [ADP-ribose] polymerase 1 | 0.367 | Evidence | |||||
| MCM2 | RRM2 | RRM2, RR2: Ribonucleoside-diphosphate reductase subunit M2 | 0.367 | Evidence | |||||
| MCM2 | PIP5K3 | PIP5K3, KIAA0981, PIKFYVE: FYVE finger-containing phosphoinositide kinase | 0.323 | Evidence | |||||
| MCM2 | YARS | YARS: Tyrosyl-tRNA synthetase, cytoplasmic | 0.323 | Evidence | |||||
| MCM2 | EIF2B5 | EIF2B5, EIF2BE: Translation initiation factor eIF-2B subunit epsilon | 0.323 | Evidence | |||||
| MCM2 | MTA1 | MTA1: Metastasis-associated protein MTA1 | 0.323 | Evidence | |||||
| MCM2 | NEO1 | NEO1, NGN: Neogenin precursor | 0.318 | Evidence | |||||
| MCM2 | ESPL1 | N/A | 0.265 | Evidence | |||||
| MCM2 | CENPA | CENPA: Histone H3-like centromeric protein A | 0.265 | Evidence | |||||
| MCM2 | MKI67 | MKI67: Antigen KI-67 | 0.265 | Evidence | |||||
| MCM2 | EIF5AP1 | EIF5A: Eukaryotic translation initiation factor 5A-1 | 0.265 | Evidence | |||||
| MCM2 | ELF1 | ELF1: ETS-related transcription factor Elf-1 | 0.265 | Evidence | |||||
| MCM2 | PSMC1 | PSMC1: 26S protease regulatory subunit 4 | 0.263 | Evidence | |||||
| MCM2 | PSMC2 | PSMC2, MSS1: 26S protease regulatory subunit 7 | 0.263 | Evidence | |||||
| MCM2 | TRIP13 | TRIP13: Thyroid receptor-interacting protein 13 | 0.263 | Evidence | |||||
| MCM2 | CBX3 | CBX3: Chromobox protein homolog 3 | 0.237 | Evidence | |||||
| MCM2 | BARD1 | BARD1: BRCA1-associated RING domain protein 1 | 0.237 | Evidence | |||||
| MCM2 | NUP153 | N/A | 0.237 | Evidence | |||||
| MCM2 | SSRP1 | SSRP1, FACT80: FACT complex subunit SSRP1 | 0.237 | Evidence | |||||
| MCM2 | SMC1L1 | SMC1A, DXS423E, KIAA0178, SB1.8, SMC1, SMC1L1: Structural maintenance of chromos | 0.237 | Evidence | |||||
| MCM2 | CDC20 | CDC20: Cell division cycle protein 20 homolog | 0.237 | Evidence | |||||
| MCM2 | KIF11 | N/A | 0.237 | Evidence | |||||
| MCM2 | LMNB1 | LMNB1, LMN2, LMNB: Lamin-B1 | 0.237 | Evidence | |||||
| MCM2 | CSPG6 | SMC3, BAM, BMH, CSPG6, SMC3L1: Structural maintenance of chromosomes protein 3 | 0.237 | Evidence | |||||
| MCM2 | LIG1 | LIG1: DNA ligase 1 | 0.237 | Evidence | |||||
| MCM2 | POLA2 | POLA2: DNA polymerase subunit alpha B | 0.237 | Evidence | |||||
| MCM2 | H2AFX | H2AFX, H2AX: Histone H2A.x | 0.237 | Evidence | |||||
| MCM2 | ANP32B | ANP32B, APRIL, PHAPI2: Acidic leucine-rich nuclear phosphoprotein 32 family memb | 0.237 | Evidence | |||||
| MCM2 | MYBL2 | MYBL2, BMYB: Myb-related protein B | 0.237 | Evidence | |||||
| MCM2 | SNRPD1 | SNRPD1: Small nuclear ribonucleoprotein Sm D1 | 0.237 | Evidence | |||||
| MCM2 | ETV6 | ETV6, TEL, TEL1: Transcription factor ETV6 | 0.237 | Evidence | |||||
| MCM2 | FOXO3A | FOXO3A, FKHRL1: Forkhead box protein O3A | 0.237 | Evidence | |||||
| MCM2 | MDN1 | MDN1, KIAA0301: Midasin | 0.237 | Evidence | |||||
| MCM2 | EXOSC2 | EXOSC2, RRP4: Exosome complex exonuclease RRP4 | 0.237 | Evidence | |||||
| MCM2 | RBMX | RBMX, HNRPG, RBMXP1: Heterogeneous nuclear ribonucleoprotein G | 0.233 | Evidence | |||||
| MCM2 | CCNB2 | CCNB2: G2/mitotic-specific cyclin-B2 | 0.228 | Evidence |

 O
O
|
||||
| MCM2 | TBP | TBP, GTF2D1, TF2D, TFIID: TATA-box-binding protein | 0.225 | Evidence |
 O
O
|
||||
| MCM2 | DDX21 | DDX21: Nucleolar RNA helicase 2 | 0.225 | Evidence | |||||
| MCM2 | DDX46 | DDX46, KIAA0801: Probable ATP-dependent RNA helicase DDX46 | 0.225 | Evidence | |||||
| MCM2 | CHD4 | CHD4: Chromodomain-helicase-DNA-binding protein 4 | 0.225 | Evidence | |||||
| MCM2 | GTF2F2 | N/A | 0.217 | Evidence | |||||
| MCM2 | ERG | ERG: Transcriptional regulator ERG | 0.217 | Evidence | |||||
| MCM2 | H1FX | H1FX: Histone H1x | 0.217 | Evidence | |||||
| MCM2 | FOXJ2 | FOXJ2, FHX: Forkhead box protein J2 | 0.217 | Evidence | |||||
| MCM2 | FXR1 | FXR1: Fragile X mental retardation syndrome-related protein 1 | 0.212 | Evidence | |||||
| MCM2 | SFRS10 | SFRS10, TRA2B: Splicing factor, arginine/serine-rich 10 | 0.21 | Evidence | |||||
| MCM2 | HNRPF | HNRPF: Heterogeneous nuclear ribonucleoprotein F | 0.21 | Evidence | |||||
| MCM2 | ELAVL1 | ELAVL1, HUR: ELAV-like protein 1 | 0.21 | Evidence | |||||
| MCM2 | CSNK2A1 | CSNK2A1: CSNK2A1 protein | 0.21 | Evidence | |||||
| MCM2 | SYNCRIP | SYNCRIP, HNRPQ, NSAP1: Heterogeneous nuclear ribonucleoprotein Q | 0.21 | Evidence | |||||
| MCM2 | EIF3S9 | EIF3S9: Eukaryotic translation initiation factor 3 subunit 9 | 0.21 | Evidence | |||||
| MCM2 | SSB | SSB: Lupus La protein | 0.21 | Evidence | |||||
| MCM2 | SFRS2 | SFRS2: Splicing factor, arginine/serine-rich 2 | 0.21 | Evidence | |||||
| MCM2 | OXSR1 | OXSR1, KIAA1101, OSR1: Serine/threonine-protein kinase OSR1 | 0.21 | Evidence | |||||
| MCM2 | SMARCA5 | SMARCA5, SNF2H, WCRF135: SWI/SNF-related matrix-associated actin-dependent regul | 0.203 | Evidence | |||||
| MCM2 | KIAA0788 | ASCC3L1, HELIC2, KIAA0788: U5 small nuclear ribonucleoprotein 200 kDa helicase | 0.203 | Evidence | |||||
| MCM2 | KHDRBS1 | KHDRBS1, SAM68: KH domain-containing, RNA-binding, signal transduction-associate | 0.203 | Evidence | |||||
| MCM2 | HSF1 | HSF1, HSTF1: Heat shock factor protein 1 | 0.195 | Evidence | |||||
| MCM2 | RFC5 | RFC5: Replication factor C subunit 5 | 0.18 | Evidence | |||||
| MCM2 | RFC4 | RFC4: Replication factor C subunit 4 | 0.18 | Evidence | |||||
| MCM2 | RFC3 | RFC3: Replication factor C subunit 3 | 0.18 | Evidence | |||||
| MCM2 | ERF | ERF: ETS domain-containing transcription factor ERF | 0.165 | Evidence | |||||
| MCM2 | IRF3 | IRF3: Interferon regulatory factor 3 | 0.165 | Evidence | |||||
| MCM2 | SMARCA3 | HLTF, HIP116A, RNF80, SMARCA3, SNF2L3, ZBU1: Helicase-like transcription factor | 0.163 | Evidence | |||||
| MCM2 | BTAF1 | BTAF1, TAF172: TATA-binding protein-associated factor 172 | 0.163 | Evidence | |||||
| MCM2 | G3BP | G3BP1, G3BP: Ras GTPase-activating protein-binding protein 1 | 0.16 | Evidence | |||||
| MCM2 | HIST1H1E | HIST1H1E, H1F4: Histone H1.4 | 0.147 | Evidence | |||||
| MCM2 | HSF2 | HSF2, HSTF2: Heat shock factor protein 2 | 0.147 | Evidence | |||||
| MCM2 | CCNL1 | CCNL1, BM-001, UNQ530/PRO1073: Cyclin-L1 | 0.147 | Evidence | |||||
| MCM2 | HIST1H1D | HIST1H1D, H1F3: Histone H1.3 | 0.147 | Evidence | |||||
| MCM2 | SRPR | SRPR: Uncharacterized protein SRPR | 0.142 | Evidence | |||||
| MCM2 | ELF2 | ELF2, NERF: ETS-related transcription factor Elf-2 | 0.142 | Evidence | |||||
| MCM2 | HTATIP | HTATIP, TIP60: Histone acetyltransferase HTATIP | 0.142 | Evidence | |||||
| MCM2 | FOXM1 | FOXM1, FKHL16, HFH11, MPP2, WIN: Forkhead box protein M1 | 0.142 | Evidence | |||||
| MCM2 | RBM14 | RBM14, SIP: RNA-binding protein 14 | 0.14 | Evidence | |||||
| MCM2 | HNRPA0 | HNRPA0: Heterogeneous nuclear ribonucleoprotein A0 | 0.14 | Evidence | |||||
| MCM2 | POLR2H | POLR2H: DNA-directed RNA polymerases I, II, and III subunit RPABC3 | 0.135 | Evidence | |||||
| MCM2 | ETS2 | ETS2: Protein C-ets-2 | 0.133 | Evidence | |||||
| MCM2 | MTIF2 | MTIF2: Translation initiation factor IF-2, mitochondrial precursor | 0.13 | Evidence | |||||
| MCM2 | HIST1H1C | HIST1H1C, H1F2: Histone H1.2 | 0.128 | Evidence | |||||
| MCM2 | CCNE1 | CCNE1, CCNE: G1/S-specific cyclin-E1 | 0.128 | Evidence | |||||
| MCM2 | CARHSP1 | CARHSP1: Calcium-regulated heat stable protein 1 | 0.128 | Evidence | |||||
| MCM2 | DHX8 | DHX8, DDX8: ATP-dependent RNA helicase DHX8 | 0.128 | Evidence | |||||
| MCM2 | RBL1 | RBL1: Retinoblastoma-like protein 1 | 0.128 | Evidence | |||||
| MCM2 | CCNT2 | CCNT2: Cyclin-T2 | 0.128 | Evidence | |||||
| MCM2 | C10orf9 | CCNY, C10orf9, CBCP1, CFP1: Cyclin-Y | 0.128 | Evidence | |||||
| MCM2 | FARSLA | FARSA, FARS, FARSL, FARSLA: Phenylalanyl-tRNA synthetase alpha chain | 0.128 | Evidence | |||||
| MCM2 | WEE1 | WEE1: Wee1-like protein kinase | 0.128 | Evidence | |||||
| MCM2 | TIAL1 | TIAL1: Nucleolysin TIAR | 0.128 | Evidence | |||||
| MCM2 | RBM15 | RBM15, OTT: Putative RNA-binding protein 15 | 0.128 | Evidence | |||||
| MCM2 | SNRPB2 | SNRPB2: U2 small nuclear ribonucleoprotein B'' | 0.128 | Evidence | |||||
| MCM2 | PABPC4 | PABPC4, APP1, PABP4: Polyadenylate-binding protein 4 | 0.128 | Evidence | |||||
| MCM2 | PABPN1 | PABPN1, PAB2, PABP2: Polyadenylate-binding protein 2 | 0.128 | Evidence | |||||
| MCM2 | PKMYT1 | PKMYT1, MYT1: Membrane-associated tyrosine- and threonine-specific cdc2-inhibito | 0.128 | Evidence | |||||
| MCM2 | SFRS4 | SFRS4, SRP75: Splicing factor, arginine/serine-rich 4 | 0.128 | Evidence | |||||
| MCM2 | PBK | PBK, TOPK: Lymphokine-activated killer T-cell-originated protein kinase | 0.128 | Evidence | |||||